Invasion Potential Alignment Alternative (Median)
vignette-026-quadrant-plots-alt-median.RmdBuild and Save Risk Correlation Quadrant Plots (Ensemble Extracts, Alternate Extraction Data)
1. Load required packages and set the same settings
# Make transport scaled between one and zero
one_zero <- TRUE
# switch between present or future transport
present_transport <- TRUE
# Set what type of multiple correlation to calculate
method_cor <- "spearman"
# set whether to look at grape or wine production (PLOT LABELS ARE NOT CLEAN FOR WINE!!!)
#impact_prod <- "wine"
impact_prod <- "grapes"
# Plot the native countries?
native <- TRUE
# remove USA?
rem_usa <- TRUE
# Replace this with max mean quantile_0.50 quantile_0.75 quantile_0.90 to edit which metric to plot for establishment
#estab_to_plot <- "obs_max" # max was used for the quadrant figures
#estab_to_plot <- "obs_mean"
estab_to_plot <- "quantile_0.50" #median2. Load the full suitability extract ensemble data and
the summary_future data (states and
countries).
Join the two dataframes and resolve conflicts. Use the
summary_future data to lead the merging with
left_join().
- mean suitability
- median suitability
#suitability data load
data("states_extracts_ensemble")
data("countries_extracts_ensemble")
#summary data load
data("states_summary_present_ensemble")
data("countries_summary_present_ensemble")
#join the data
states_summary_present_ensemble <- states_summary_present_ensemble %>%
left_join(., states_extracts_ensemble, by = "geopol_unit") %>%
#drop grandmeanmax col, it is redundant
dplyr::select(-grand_mean_max)
countries_summary_present_ensemble <- countries_summary_present_ensemble %>%
left_join(., countries_extracts_ensemble, by = "geopol_unit") %>%
#drop grandmeanmax col, it is redundant
dplyr::select(-grand_mean_max)
#make it easier to port over parts of the vig-021 code by changing the names
states_summary_present <- states_summary_present_ensemble
countries_summary_present <- countries_summary_present_ensemble3. Drop the same states and countries that
we dropped in the regular correlation plot.
#filter out some No Data for ensemble model
countries_summary_present <- countries_summary_present %>%
filter(!ID %in% c("Antarctica", "Monaco", "Norfolk Island", "Spratly Islands"))
if(rem_usa){ # TRUE if you want to remove the USA
rems <- c("Akrotiri and Dhekelia"
,"?land"
,"American Samoa"
,"Bouvet Island"
,"British Indian Ocean Territory"
,"Caspian Sea"
,"Christmas Island"
,"Clipperton Island"
,"Cocos Islands"
,"Falkland Islands"
,"Faroe Islands"
,"French Southern Territories"
,"Heard Island and McDonald Islands"
,"Mayotte"
,"Northern Mariana Islands"
,"Paracel Islands"
,"Pitcairn Islands"
,"Saint Pierre and Miquelon"
,"Tokelau"
,"United States Minor Outlying Islands"
,"Wallis and Futuna"
,"United States"
,"Philippines")
} else {
rems <- c("Akrotiri and Dhekelia"
,"?land"
,"American Samoa"
,"Bouvet Island"
,"British Indian Ocean Territory"
,"Caspian Sea"
,"Christmas Island"
,"Clipperton Island"
,"Cocos Islands"
,"Falkland Islands"
,"Faroe Islands"
,"French Southern Territories"
,"Heard Island and McDonald Islands"
,"Mayotte"
,"Northern Mariana Islands"
,"Paracel Islands"
,"Pitcairn Islands"
,"Saint Pierre and Miquelon"
,"Tokelau"
,"United States Minor Outlying Islands"
,"Wallis and Futuna"
,"Philippines")
}
#do the rm now
countries_summary_present <- countries_summary_present %>% filter(!ID %in% rems)
#set the native countries
if(native){
countries_summary_present$status[countries_summary_present$geopol_unit %in% c("China", "India", "Taiwan", "Vietnam")] <- "native"
} else {
countries_summary_present$status[countries_summary_present$geopol_unit %in% c("China", "India", "Taiwan", "Vietnam")] <- "not established"
}4. Rescale the data the same way but also select the metric to look at.
# scale the import data
if(one_zero){
states_summary_present <- states_summary_present %>%
# transport
mutate(avg_infected_mass_scaled = log10(avg_infected_mass+1)) %>%
mutate(avg_infected_mass_scaled = (avg_infected_mass_scaled - min(avg_infected_mass_scaled))) %>%
mutate(avg_infected_mass_scaled = avg_infected_mass_scaled / max(avg_infected_mass_scaled)) %>%
# establishment
#mutate(suitability_scaled = .[[estab_to_plot]])
#mutate(suitability_scaled = .[[estab_to_plot]] / max(.[[estab_to_plot]]))
mutate(suitability_scaled = .[[estab_to_plot]] - min(.[[estab_to_plot]])) %>%
mutate(suitability_scaled = (suitability_scaled / max(suitability_scaled)))
} else {
states_summary_present <- states_summary_present %>%
# transport
mutate(avg_infected_mass_scaled = log(avg_infected_mass+1)) %>%
# establishment
mutate(suitability_scaled = .[[estab_to_plot]])
}
if(one_zero){
countries_summary_present <- countries_summary_present %>%
# transport
mutate(avg_infected_mass_scaled = log10(avg_infected_mass+1)) %>%
mutate(avg_infected_mass_scaled = (avg_infected_mass_scaled - min(avg_infected_mass_scaled))) %>%
mutate(avg_infected_mass_scaled = avg_infected_mass_scaled / max(avg_infected_mass_scaled)) %>%
# establishment
#mutate(suitability_scaled = .[[estab_to_plot]])
#mutate(suitability_scaled = .[[estab_to_plot]] / max(.[[estab_to_plot]]))
mutate(suitability_scaled = .[[estab_to_plot]] - min(.[[estab_to_plot]])) %>%
mutate(suitability_scaled = (suitability_scaled / max(suitability_scaled)))
} else {
countries_summary_present <- countries_summary_present %>%
# transport
mutate(avg_infected_mass_scaled = log(avg_infected_mass+1)) %>%
# establishment
mutate(suitability_scaled = .[[estab_to_plot]])
}5. Plotting settings
# label both of the countries (or states) with established SLF?
label.countries.est <- TRUE
label.states.est <- TRUE #same for both present/future: CT, DE, MD, NJ, OH, WV
# labeling aesthetics for label.countries.est and label.states.est
nudge_countries.est.x <- c(.05,0.05)
nudge_countries.est.y <- c(.12,0.11)
nudge_countries.est_future.x <- c(.05,0.05) #jpn and kor
nudge_countries.est_future.y <- c(.12,0.11)
nudge_states.est.x <- c( 0.00, 0.00, -0.10, 0.02, 0.01, 0.00) #adding OH, CT, DE, MD, NJ, WV
nudge_states.est.y <- c(-0.10, -0.10, 0.10, -0.17,-0.15,-0.10)
# Choose quadrant intercepts
q_intercepts <- list(transport = 0.5, establishment = 0.5)
#set margins
hh = .001
# Do not change this flips between plotting the states or plotting countries
plots_which <- c(TRUE,FALSE)
# How many countries and states to highlight for labels?
top_to_plot <- 10
#color aesthetics
risk_color <- "gray40"
risk_size <- 5
#lx <- 0.22
#ly <- 0.25
#hy <- 0.77
#hx <- 0.78
#
#risk_labels <-data.frame(label = c("low\nrisk","moderate\nrisk","moderate\nrisk","high\nrisk"),
# x = c(lx,lx,hx,hx),
# y = c(ly,hy,ly,hy),
# hjust = c(0,0,0,0),
# vjust = c(0,0,0,0))
#constant looping var
i <- 16. Plotting code itself
for (i in 1:length(plots_which)) {
#selects states or countries to plot
statez = plots_which[i]
########################
#DATA SELECTION STEP
########################
#checks whether to use wine or grapes as the impact product to consider
if(impact_prod == "wine"){
if (statez) {
data_to_plot <- states_summary_present
#calls the exact variables to plot
data_to_plot <- data_to_plot %>%
mutate(
x_to_plot = avg_infected_mass_scaled,
y_to_plot = suitability_scaled,
fill_to_plot = grapes,
size_to_plot = avg_wine,
color_to_plot = status
) %>%
arrange(desc(grapes), (size_to_plot)) #so that the grape producers are on top
#order the data that gets labeled based on top_to_plot
data_to_label <- tail(data_to_plot, top_to_plot) %>% arrange(desc(size_to_plot))
#do welch t.test to assess if transport and establishment have the same relationship with grapes
#state_grape_t.test <- list(transport = t.test(avg_infected_mass_scaled~grapes, data = data_to_plot),
# establishment = t.test(suitability_scaled~grapes, data = data_to_plot))
#add infected state labeling if turned on
if(label.states.est){
#PRESENT
data_to_label_states.est <- data_to_plot %>%
filter(geopol_unit %in% c("Connecticut", "Delaware", "Maryland", "New Jersey", "Ohio", "West Virginia")) %>%
arrange(desc(size_to_plot))
data_to_label <- bind_rows(data_to_label,data_to_label_states.est)
} #end of labeling for infected states
} else {
#selects the data to modify for plotting: COUNTRIES and selects the timing
data_to_plot <- countries_summary_present
data_to_plot <- data_to_plot %>%
mutate(
x_to_plot = avg_infected_mass_scaled,
y_to_plot = suitability_scaled,
fill_to_plot = grapes,
size_to_plot = avg_wine,
color_to_plot = status
) %>%
arrange(desc(grapes), (size_to_plot)) #so that the grape producers are on top
data_to_label <-
tail(data_to_plot, top_to_plot) %>%
arrange(desc(size_to_plot)) %>%
mutate(
ID = recode( #add ISO3 codes for the labeled countries (rather than all)
ID,
Italy = "ITA",
France = "FRA",
Spain = "ESP",
China = "CHN",
Argentina = "ARG",
Chile = "CHL",
Australia = "AUS",
`South Africa` = "ZAF",
Germany = "DEU",
Portugal = "PRT"
)
)
#do welch t.test to assess if transport and establishment have the same relationship with grapes
#country_grape_t.test <- list(transport = t.test(avg_infected_mass_scaled~grapes, data = data_to_plot),
# establishment = t.test(suitability_scaled~grapes, data = data_to_plot))
if(label.countries.est){
data_to_label_countries.est <- data_to_plot %>%
filter(geopol_unit %in% c("Japan", "South Korea")) %>%
mutate(ID = recode(ID,
Japan = "JPN",
`South Korea` = "KOR")) %>%
arrange(desc(size_to_plot))
data_to_label <- bind_rows(data_to_label,data_to_label_countries.est)
}
}
} else if(impact_prod == "grapes"){
if (statez) {
#selects the data to modify for plotting: STATES and chooses the timing
data_to_plot <- states_summary_present
data_to_plot <- data_to_plot %>%
mutate(
x_to_plot = avg_infected_mass_scaled,
y_to_plot = suitability_scaled,
fill_to_plot = wine,
size_to_plot = avg_prod, #avg_yield or avg_prod are the two options here for grapes
color_to_plot = status
) %>%
arrange(desc(grapes), (size_to_plot)) #so that the grape producers are on top
#change the zeros to one's for plot size
data_to_plot$size_to_plot[data_to_plot$size_to_plot == 0] <- 1
data_to_label <-
tail(data_to_plot, top_to_plot) %>% arrange(desc(size_to_plot))
#state_grape_t.test <- list(transport = t.test(avg_infected_mass_scaled~grapes, data = data_to_plot),
# establishment = t.test(suitability_scaled~grapes, data = data_to_plot))
#add infected state labeling if turned on
if(label.states.est){
#PRESENT
data_to_label_states.est <- data_to_plot %>%
filter(geopol_unit %in% c("Connecticut", "Delaware", "Maryland", "New Jersey", "Ohio", "West Virginia")) %>%
arrange(desc(size_to_plot))
data_to_label <- bind_rows(data_to_label,data_to_label_states.est)
} #end of labeling for infected states
} else {
#selects the data to modify for plotting: COUNTRIES and selects the timing
data_to_plot <- countries_summary_present
data_to_plot <- data_to_plot %>%
mutate(
x_to_plot = avg_infected_mass_scaled,
y_to_plot = suitability_scaled,
fill_to_plot = wine,
size_to_plot = avg_prod, #avg_yield or avg_prod are the two options here for grapes
color_to_plot = status
) %>%
arrange(desc(grapes), (size_to_plot)) #so that the grape producers are on top
#change the zeros to one's for plot size
data_to_plot$size_to_plot[data_to_plot$size_to_plot == 0] <- 1
data_to_label <-
tail(data_to_plot, top_to_plot) %>% arrange(desc(size_to_plot))
data_to_label <- data_to_label %>%
mutate(
ID = recode(
ID,
Egypt = "EGY",
Peru = "PER",
India = "IND",
Albania = "ALB",
Iraq = "IRQ",
Brazil = "BRA",
Thailand = "THA",
`South Africa` = "ZAF",
China = "CHN",
Armenia = "ARM",
Italy = "ITA",
Spain = "ESP",
France = "FRA",
Turkey = "TUR",
Chile = "CHL",
Argentina = "ARG",
Australia = "AUS",
`South Korea` = "KOR",
Japan = "JPN"
)
)
# country_grape_t.test <- list(transport = t.test(avg_infected_mass_scaled~grapes, data = data_to_plot),
# establishment = t.test(suitability_scaled~grapes, data = data_to_plot))
if(label.countries.est){
data_to_label_countries.est <- data_to_plot %>%
filter(geopol_unit %in% c("Japan", "South Korea")) %>%
mutate(ID = recode(ID,
Japan = "JPN",
`South Korea` = "KOR")) %>%
arrange(desc(size_to_plot))
data_to_label <- bind_rows(data_to_label,data_to_label_countries.est)
} # end of if label plotting JPN KOR
}
}
########################
#PLOT SETTINGS STEP
########################
#SET ALL OF THE NUDGES FOR LABELS
#need to add the conditional for present transport, since nudging only works based on the data
if(present_transport == FALSE){
if(impact_prod == "wine"){
#nudging
if(statez) { #STATES FUTURE WINE
nudge.x <- c(-.1,-.1,-.1,-.05,-.1,-0.05,-.1,.06,-.01,.05)
nudge.y <- c(.2,.10,.10,.10,.07,.15,.15,-.07,-.12,-.12) #CA, WA, NY, PA, OR, GA, OH, MI, VA, NC
if(label.states.est){
nudge.x <- c(nudge.x, nudge_states.est.x)
nudge.y <- c(nudge.y, nudge_states.est.y)
}
ytitle <- "Establishment Potential"
} else { #COUNTRIES FUTURE WINE
nudge.x <- c(-.3,-.27,-.05,-.26,.14,.1,.20,-.21,.08,.2)
nudge.y <- c(.13, .095, .155, .09, -.11, .13, .15,.06,-.1,-.19) #ITA, FRA, ESP, CHN, ARG, CHL, AUS, ZAF, DEU, PRT
if(label.countries.est) {
nudge.x <- c(nudge.x, nudge_countries.est_future.x) #JPN KOR
nudge.y <- c(nudge.y, nudge_countries.est_future.y) #JPN KOR
}
ytitle <- "Establishment Potential"
}
} else if(impact_prod == "grapes"){
#nudging
if(statez) { #STATES FUTURE GRAPES
nudge.x <- c(-.00,-.05,-.06,-.02, .10,-.05,-.01,-.01,-.05,-.03) #yield: #CA, PA, WA, MI, NY, MO, OH, OR, GA, NC
nudge.y <- c(-.10, .10, .08, .08,-.05, .09,-.10,-.10, .09, .10) #production: CA, WA, NY, PA, MI, OR, TX, VA, NC, MO
if(label.states.est){
nudge.x <- c(nudge.x, nudge_states.est.x)
nudge.y <- c(nudge.y, nudge_states.est.y)
}
ytitle <- "Establishment Potential"
} else { #COUNTRIES FUTURE GRAPES
#ensemble version
# grape prod: CHN, ITA, ESP, FRA, TUR, IND, CHL, ARG, ZAF, AUS
nudge.x <- c(-0.00,-0.00,-0.01,-0.02,-0.00, 0.03,-0.02, 0.03,-0.02,-0.00)
nudge.y <- c(-0.10, 0.085, 0.15, 0.07,-0.10, 0.05, 0.13,-0.15, 0.29,-0.12)
#old version
# grape production: CHN, ITA, ESP, FRA, TUR, IND, CHL, ARG, ZAF, AUS
#nudge.x <- c(-0.01,-0.00,-0.00,-0.04, 0.02,0.03,-0.04, 0.05,-0.03, 0.04)
#nudge.y <- c(-0.10,-0.20,-0.27, 0.06,-0.24,0.07, 0.11,-0.23, 0.16,-0.10)
if(label.countries.est) {
nudge.x <- c(nudge.x, nudge_countries.est_future.x) #JPN KOR
nudge.y <- c(nudge.y, nudge_countries.est_future.y) #JPN KOR
}
ytitle <- "Establishment Potential"
}
}
} else { # grapes above wine below NOTE may not work with labeling JPN and KOR
if(impact_prod == "wine"){
if(statez) { #STATES PRESENT WINE
nudge.x <- c(.09,-.1,-.04,-.05,-.10,-0.12,-.1,.04,-.01,.05)
nudge.y <- c(-.10,.10,.10,.10,.07,.10,.12,-.07,-.12,-.12) #CA, WA, NY, PA, OR, GA, OH, MI, VA, NC
if(label.states.est){
nudge.x <- c(nudge.x, nudge_states.est.x)
nudge.y <- c(nudge.y, nudge_states.est.y)
}
ytitle <- "Establishment Potential"
} else { #COUNTRIES PRESENT WINE
nudge.x <- c(-.03,-.10,-.02,-.03,.14,-.1,-.23,-.21,.08,.2)
nudge.y <- c(.13, .09, .155, .09, -.11, .13, .15,.09,-.08,-.15) #ITA, FRA, ESP, CHN, ARG, CHL, AUS, ZAF, DEU, PRT
if(label.countries.est) {
nudge.x <- c(nudge.x, nudge_countries.est.x) #JPN KOR
nudge.y <- c(nudge.y, nudge_countries.est.y) #JPN KOR
}
ytitle <- "Establishment Potential"
}
} else if(impact_prod == "grapes"){
if(statez) { #STATES PRESENT GRAPES
nudge.x <- c(-.00,-.05,-.06,-.02, .10,-.05,-.01,-.01,-.05,-.03) #yield: #CA, PA, WA, MI, NY, MO, OH, OR, GA, NC
nudge.y <- c(-.10, .10, .08, .08,-.05, .09,-.10,-.10, .09, .10) #production: CA, WA, NY, PA, MI, OR, TX, VA, NC, MO
if(label.states.est){
nudge.x <- c(nudge.x, nudge_states.est.x)
nudge.y <- c(nudge.y, nudge_states.est.y)
}
ytitle <- "Establishment Potential"
} else { #COUNTRIES PRESENT GRAPES
#ensemble version
# grape prod: CHN, ITA, ESP, FRA, TUR, IND, CHL, ARG, ZAF, AUS
nudge.x <- c(-0.00,-0.00,-0.01,-0.02,-0.00, 0.03,-0.02, 0.05,-0.02, 0.01)
nudge.y <- c(-0.10, 0.08, 0.15, 0.07,-0.10, 0.05, 0.13,-0.12, 0.29,-0.09)
#nudge.x <- c(-0.01, 0.02, 0.10,-0.04, 0.06,-0.02,-0.07, 0.05,-0.20, 0.11) #CHN, ITA, ESP, FRA, TUR, IND, CHL, ARG, ZAF, AUS
#nudge.y <- c( 0.08,-0.08,-0.20, 0.07,-0.02, 0.10, 0.12,-0.09,-0.07,-0.25)
if(label.countries.est) {
nudge.x <- c(nudge.x, nudge_countries.est.x) #JPN KOR
nudge.y <- c(nudge.y, nudge_countries.est.y) #JPN KOR
}
ytitle <- "Establishment Potential"
}
}
}
#CREATE THE STATES PLOT
(states_plot <- ggplot(data = data_to_plot) +
#geom_text(data = risk_labels, mapping = aes(x = x, y = y,label = label), color = risk_color, size = risk_size) +
geom_hline(yintercept = q_intercepts$establishment, linetype = "dashed") +
geom_vline(xintercept = q_intercepts$transport, linetype = "dashed") +
# geom_abline(intercept = 0, slope = 1, linetype = "solid") + # add one to one
geom_rect(mapping = aes(xmin=1.01, xmax=1.2, ymin=.49, ymax=.5), fill = "white") +
geom_rect(mapping = aes(ymin=1.01, ymax=1.2, xmin=.49, xmax=.51), fill = "white") +
geom_text_repel(data = data_to_label,
aes(x = x_to_plot, y = y_to_plot, label = ID),
min.segment.length = 0,
direction = "x",
nudge_y = nudge.y,
nudge_x = nudge.x
) +
geom_point(
aes(x = x_to_plot, y = y_to_plot, fill = fill_to_plot, size = size_to_plot, color = color_to_plot),
shape = 21, stroke = 1.3, alpha = 0.75
) +
scale_fill_manual(
values = c("no" = "#ffffff", "yes" = "#C77CFF"),
name = "Wine Production",
labels = c("high", "low")
) +
scale_color_manual(
values = c(
"red",
"black",
"blue"),
breaks = c(
"established",
"not established",
"native"),
name = "Regional Status",
labels = c("invaded","uninvaded", "native")
) +
guides(
fill = guide_legend(
order = 1,
override.aes = list(shape = 22, size = 5, alpha = 1)
),
color = guide_legend(
order = 3,
override.aes = list(alpha = 1)
),
size = guide_legend( # Adjust size to edit legend grape production
order = 2, # circles.
override.aes = list(size = c(1,3,6), alpha = 1)
)
) +
scale_size_continuous(name = "Grape Production",
trans = "log10",
range = c(1.5, 6),
breaks = c(1.5,2,6),
labels = c("low", "moderate", "high")
) +
labs(x = "Transport Potential", y = ytitle) +
ylim(0, 1.2) + xlim(0, 1.2) +
theme(
panel.grid.major = element_line(colour = "#f0f0f0"),
panel.grid.minor = element_blank(),
#panel.grid.major = element_blank(),
#panel.border = element_rect(colour = "black", fill=NA, size=1),
axis.line = element_line(colour = "black"),
legend.position = c(0.7, 0.25),
panel.background = element_blank(),
plot.background = element_blank(),
legend.background = element_blank(),#element_rect(colour = 'black', fill = 'white', linetype='solid'),
axis.text = element_text(size = rel(1)),
axis.title.x = element_text(hjust = .4),
axis.title.y = element_text(hjust = .35),
legend.title = element_text(face = "plain"),
legend.key.size = unit(0.2, "cm"),
legend.key = element_blank(),
plot.margin = unit(c(hh, -5, hh, hh), units = "line"),
axis.title = element_text(size = rel(1.3))
) +
scale_x_continuous(breaks = c(0,.5,1),labels = c("low", "moderate", "high")) +
scale_y_continuous(breaks = c(0,.5,1),labels = c("low", "moderate", "high")) +
coord_flip()
)
#get the legend
get_legend <- function(myggplot) {
tmp <- ggplot_gtable(ggplot_build(myggplot))
leg <-
which(sapply(tmp$grobs, function(x)
x$name) == "guide-box")
legend <- tmp$grobs[[leg]]
return(legend)
}
#plot all of it
if(statez) {
legend_state <- legend <- get_legend(states_plot)
} else {
legend_country <- legend <- get_legend(states_plot)
}
(states_plot <- states_plot + theme(legend.position = "none"))
#boxplot: ESTABLISHMENT
(
states_box_estab <- ggplot(data_to_plot) +
geom_boxplot(
aes(x = fill_to_plot, y = y_to_plot, group = fill_to_plot, fill = fill_to_plot),
show.legend = FALSE,
outlier.shape = NA,
notch = FALSE,
notchwidth = .25
) +
ylim(0, 1) +
scale_fill_manual(values = c("yes" = "#C77CFF", "no" = "#ffffff"), name = "") +
theme(
axis.line = element_blank(),
axis.text.x = element_blank(),
axis.text.y = element_blank(),
axis.ticks = element_blank(),
axis.title.x = element_blank(),
axis.title.y = element_blank(),
legend.position = "none",
panel.background = element_blank(),
panel.border = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
plot.margin = unit(c(hh, hh, hh, hh), units = "line"),
plot.background = element_blank()
) +
coord_flip()
)
#boxplot: INTRODUCTION
(
states_box_intro <- ggplot(data_to_plot) +
geom_boxplot(
aes(x = fill_to_plot, y = x_to_plot, group = fill_to_plot, fill = fill_to_plot),
show.legend = FALSE,
outlier.shape = NA,
notch = FALSE,
notchwidth = .25
) +
labs(x = " ", y = NULL) +
ylim(0, 1) +
scale_fill_manual(
values = c("yes" = "#C77CFF", "no" = "#ffffff"),
name = ""
) +
theme(
axis.line = element_blank(),
axis.text.x = element_blank(),
axis.text.y = element_blank(),
axis.ticks = element_blank(),
axis.title.x = element_blank(),
legend.position = "none",
panel.background = element_blank(),
panel.border = element_blank(),
panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
plot.background = element_blank(),
plot.margin = unit(c(hh, hh, hh, hh), units = "line")
)
)
(
states_plot_final <- states_plot +
annotation_custom(
grob = ggplotGrob(states_box_estab),
xmin = 1.01,
xmax = 1.11,
ymin = -Inf,
ymax = 1.05
) +
# Insert ybp_grob inside the scatter plot
annotation_custom(
grob = ggplotGrob(states_box_intro),
xmin = -.1,
xmax = 1.05,
ymin = 1,
ymax = 1.1
) +
annotation_custom(
grob = rectGrob(gp = gpar(fill = "white", col = "white")),
xmin = -1,
xmax = 1.21,
ymin = 1.1,
ymax = 1.21
) +
annotation_custom(
grob = rectGrob(gp = gpar(fill = "white", col = "white")),
ymin = -1,
ymax = 1.21,
xmin = 1.11,
xmax = 1.21
) +
scale_x_continuous(breaks = c(0,.5,1),labels = c("low", "moderate", "high"), limits = c(0,1.15)) +
scale_y_continuous(breaks = c(0,.5,1),labels = c("low", "moderate", "high"), limits = c(0,1.15))
)
g <- ggplotGrob(states_plot_final)
g$layout$clip[g$layout$name == "panel"] <- "off"
#g$layout
g$layout$z[g$layout$name == "panel"] <- 17 # Note that z for panel is 1. Change it to something bigger.
#Correlation Analysis
# present or future
if(present_transport){
if (statez) { #states present
states_grob <- g
} else { #countries present
countries_grob <- g
}
} else { #future cases
if (statez) { #states future
states_grob <- g
} else{ #countries future
countries_grob <- g
}
}
#SAVES RESULTS AS PDF
#plot the states+legend and save
if (statez) {
#pdf(file.path(here::here(),"vignettes", paste0("states_combined_future_max_",impact_prod,".pdf")),width = 7.5, height = 7)
grid.arrange(states_grob, legend_state, nrow=1, ncol = 2, widths = c(4,2), heights = 1)
invisible(dev.off())
} else{
#plot the countries+legend
#pdf(file.path(here::here(),"vignettes", paste0("countries_combined_future_max_",impact_prod,".pdf")),width = 7.5, height = 7)
grid.arrange(countries_grob, legend_country, nrow=1, ncol = 2, widths = c(4,2), heights = 1)
invisible(dev.off())
}
} # end of the 2 loop for loop for plotting states and countries7. Display the plots
#set up a layout grid
lay1 <- rbind(c( 1, 1, 1,NA,NA,NA),
c( 1, 1, 1,NA,NA,NA),
c( 1, 1, 1,NA, 3, 3),
c(NA,NA,NA,NA, 3, 3),
c( 2, 2, 2,NA, 3, 3),
c( 2, 2, 2,NA,NA,NA),
c( 2, 2, 2,NA,NA,NA)
)
#nice version for knitting
grid.arrange(states_grob, countries_grob, legend_country, nrow = 7, ncol = 6, layout_matrix = lay1, heights = c(2,2,2, 0, 2,2,2), widths = c(1,1,1,0.1,0.6,0.6))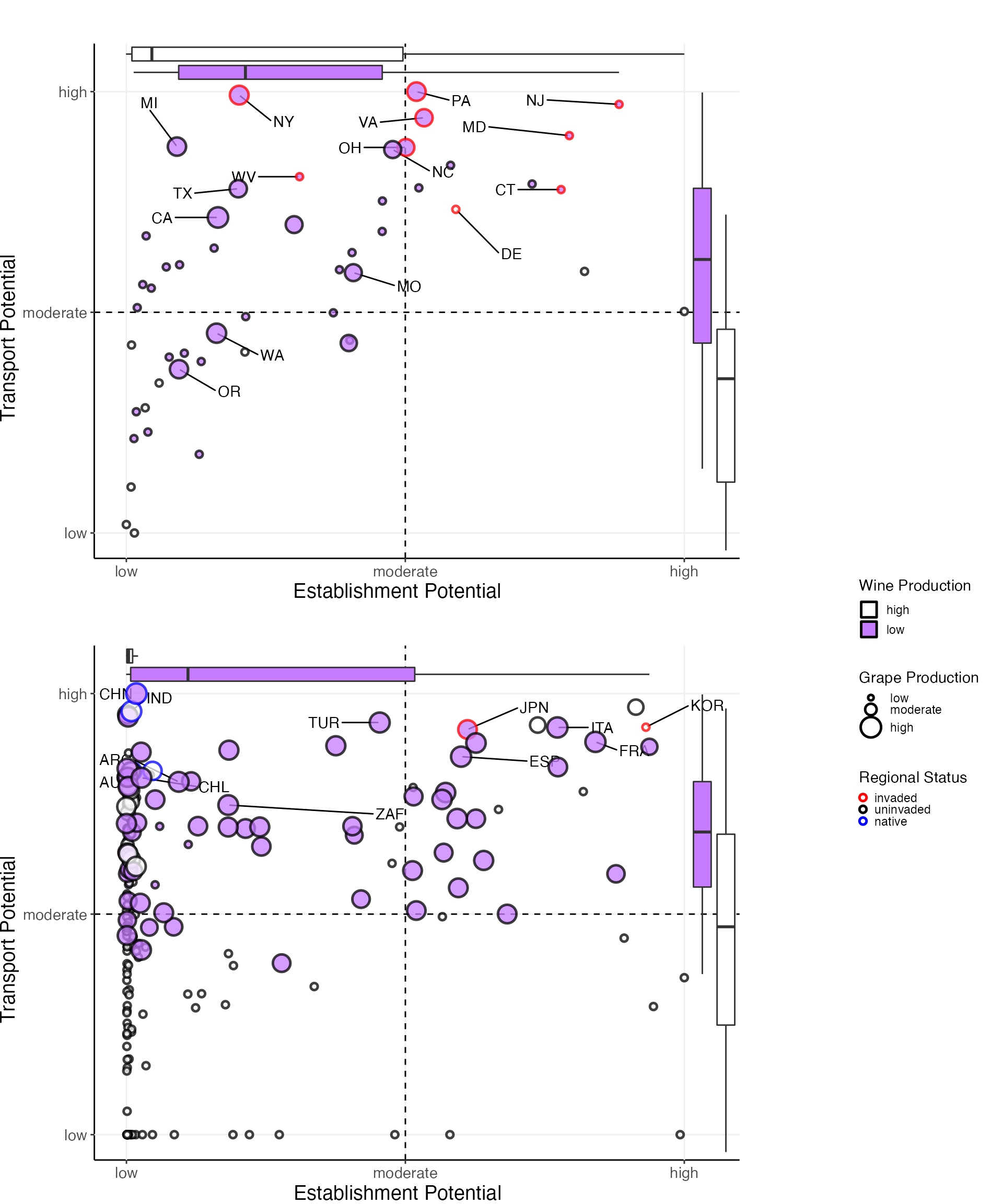
8. Get and print the correlations
#grapes
#STATES
#build model
states_grapes_impact_model <- lm(formula = log10_avg_prod ~ avg_infected_mass_scaled + suitability_scaled, data = states_summary_present)
#do cor
states_grapes_impact_cor <- cor.test(states_grapes_impact_model$model$log10_avg_prod, states_grapes_impact_model$fitted.values, method = method_cor)
#COUNTRIES
#build model
countries_grapes_impact_model <- lm(formula = log10_avg_prod ~ avg_infected_mass_scaled + suitability_scaled, data = countries_summary_present %>%
filter(!ID %in% c("United States",
"China",
"India",
"Taiwan",
"Japan",
"South Korea",
"Vietnam")))
#do cor
countries_grapes_impact_cor <- cor.test(countries_grapes_impact_model$model$log10_avg_prod, countries_grapes_impact_model$fitted.values, method = method_cor)
#wine
#STATES
#model
states_wine_impact_model <- lm(formula = log10_avg_wine ~ avg_infected_mass_scaled + suitability_scaled, data = states_summary_present)
#cor
states_wine_impact_cor <- cor.test(states_wine_impact_model$model$log10_avg_wine, states_wine_impact_model$fitted.values, method = method_cor)
#COUNTRIES
#model
countries_wine_impact_model <- lm(formula = log10_avg_wine ~ avg_infected_mass_scaled + suitability_scaled, data = countries_summary_present %>%
filter(!ID %in% c("United States",
"China",
"India",
"Taiwan",
"Japan",
"South Korea",
"Vietnam")))
#cor
countries_wine_impact_cor <- cor.test(countries_wine_impact_model$model$log10_avg_wine, countries_wine_impact_model$fitted.values, method = method_cor)We can report the following alignment of potentials:
- GRAPES
- U.S. State Alignment Correlation = 0.446, p = 1.031 x 10-03
- Country Alignment Correlation = 0.584, p = 9.622 x 10-22
- WINE
- U.S. State Alignment Correlation = 0.613, p = 1.780 x 10-06
- Country Alignment Correlation = 0.527, p = 2.585 x 10-17
Note: grapes is the plotted impact product and quantile_0.50 is the metric of interest.